Rognes, T.: Faster Smith–Waterman database searches with inter-sequence simd parallelisation. Science 349(6245), 261–266 (2015)Įnright, A.J., Ouzounis, C.A.: Generage: a robust algorithm for sequence clustering and domain detection. Hirschberg, J., Manning, C.D.: Advances in natural language processing. Krasnogor, N., Pelta, D.A.: Measuring the similarity of protein structures by means of the universal similarity metric. Pinkel, D., Albertson, D.G.: Array comparative genomic hybridization and its applications in cancer. In: 2008 IEEE Fourth International Conference on eScience, pp. The results showed that the proposed hybrid approach was up to 242 times faster than the sequential approach.įortes, J., Matsunaga, A., Tsugawa, M.: Cloudblast: combining mapreduce and virtualization on distributed resources for bioinformatics applications. Moreover, the scalability of the approach is verified on randomly generated sequences with predefined similarity levels. 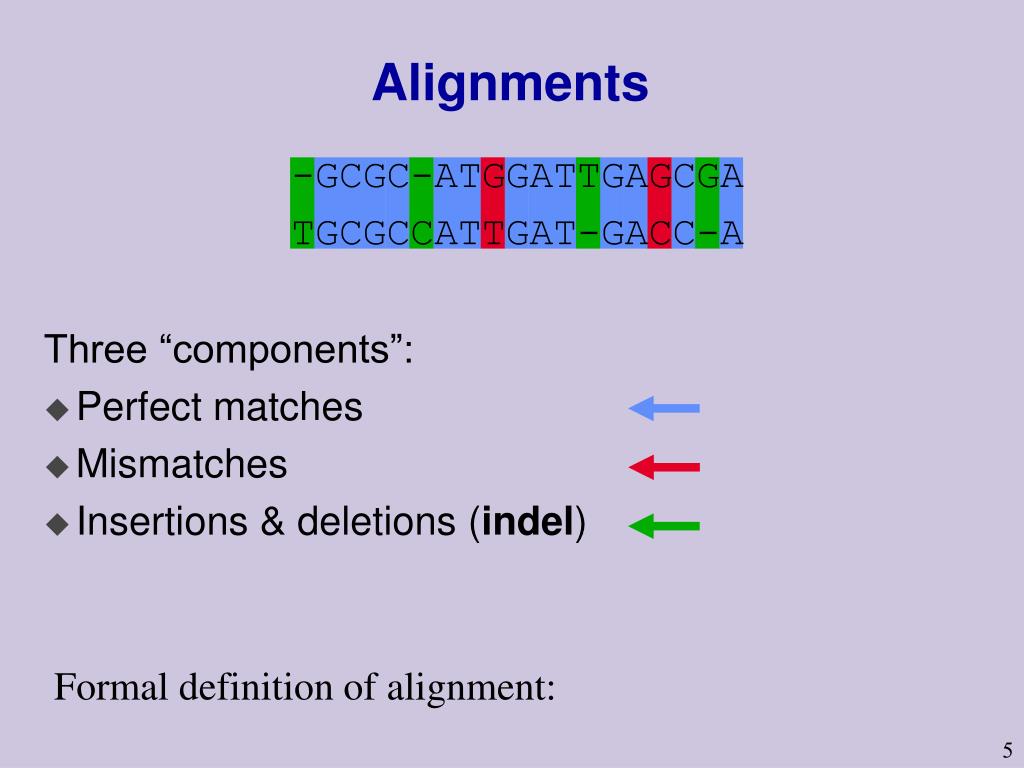
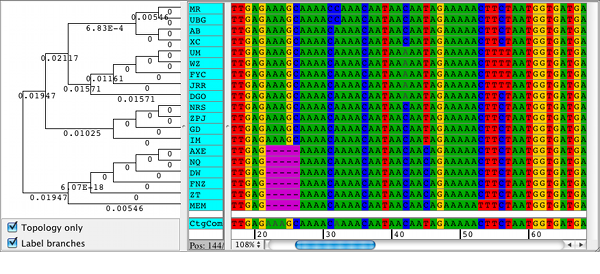
The validity of our approach is tested on real protein sequences. Thus, we propose a hybrid parallel approach that combines the capabilities of multi-core CPUs and the power of contemporary GPUs, and significantly speeds up the execution of the target algorithms. These two algorithms are computationally expensive which hinders their applicability for large data sets. In this paper, we target two well-known protein sequence alignment algorithms, the Needleman–Wunsch and the Smith–Waterman algorithms. Protein sequence alignment is concerned with identifying the similarities and the relationships among different protein structures. Understanding cellular processes contributes to the development of drugs for metabolic pathways. Protein structure and sequence analysis are vital to the understanding of cellular processes. Bioinformatics involves several computational techniques such as sequence and structural alignment, data mining, macromolecular geometry, prediction of protein structure and gene finding. Bioinformatics is an interdisciplinary field that applies trending techniques in information technology, mathematics, and statistics in studying large biological data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
